A new paper on Tibetan autosomal genetics holds some information relevant not just for Tibetans but for other East Asian and in general human peoples:
Binbin Wang et al., On the Origin of Tibetans and Their Genetic Basis in Adapting High-Altitude Environments. PLoS ONE 2011. Open access.
The generic place of Tibetans within Humankind
This is maybe the information of most generic interest, as it requires to map not just Tibetans and neighbors but also peoples from all over the world, notably Eurasia.
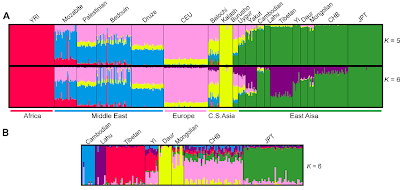
For this, figure 1 holds nearly all the relevant information. Some sub-graphs follow:
Above, the global Frappe structure results (A) place Tibetans clearly within East Asian populations (green) but show a distinctiveness of their own since K=6 (purple). In spite of sharing Y-DNA macro-haplogroup D with Japanese, Tibetans show no particular affinity with these (JPT).
In the East Asian specific results (B), at K=6, Tibetans again appear on their own, as do the other ethnic groups, except the Yi and Mongolians, who are subsumed within their neighbors to a great extent.
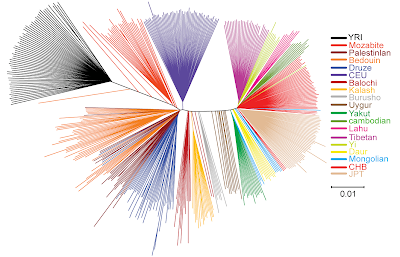
In the neighbor-joining phylogenetic tree (above), we can see that Tibetans again fall fully within the East Asian group (right fan). This structure is more interesting at this point for the global context anyhow because it can explain, for example, why in some cases West Eurasians and Africans may cluster together versus East Asians, depending on sampling conditions and probably the presence or absence of Indians (only some Pakistanis represent South Asia in this graph, as in so many others) as well as the numbers of sampled African (typically low because they are used as control only). If you draw a median line in this NJ tree, East Asians, in spite of their tight affinity with each other, fall to one half and West Eurasians and Africans fall to the other together.
Whatever this means, we can easily see that East Asians share an ancestral founder effect (including also Melanesians per previous research). However I must warn that this is more genuinely apparent in the HUGO consortium paper than here, because of the lack of Indian samples in this case. See here, here and here, for my own old entries in this matter (it is particularly clear in this graph).
By comparison only very homogeneous groups share such marked apparent founder effects, for example Europeans, West Africans, Pygmies, some (but not other) Indians. All these in the HUGO paper, in this paper instead, we can see again Europeans (but only represented by CEU, what is less meaningful), most Mozabites, most Bedouins, the other Levant populations (Palestinians, Druze and some Bedouins) and, very markedly, the Kalash.
However in the haploid genetic aspect East Asians do not seem so extremely “bottlenecked” as they do appear with autosomal genetics, not at all. In fact they host huge diversity within Eurasian haploid genetics. This is hard to explain, admittedly, but it clearly sends an alert signal for us to be extremely cautious and try to comprehend what really happened in this part of the World.
Tibetans are most closely related to the Yi
The term Yi (Nuoso in one of their languages, Lolo in Vietnam and Thailand) is a catchall term from Chinese ethnography to refer to an array of Tibeto-Burman-speaking peoples from Yunnan and nearby areas.
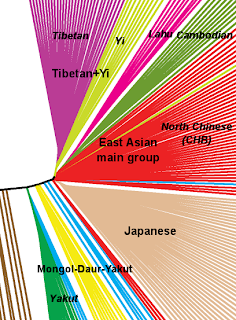
The close affinity between Tibetans and Yi in the context of East Asian populations is apparent when we apply the microscope, so to say, to the NJ tree (pictured above), as I did here:
It is easy to appreciate that, excluding the Mongol-Daur-Yakut loose group (suspect of minor West Eurasian admixture), the narrow East Asian founder effect split soon in three groups (of those shown here): Tibetans+Yi, the main group (Chinese, Cambodian and Lahu) and the Japanese – each of which diversified rapidly from those specific roots.
However this result is not coincident with the HUGO consortium tree (different method: maximum likelihood), where Cambodians fall in a SE Asian specific group and Chinese in a distinct SEA/EA/America shared one. So, as always, be careful before rushing to conclusions based only on one paper’s data.
High altitude genes have been selected among Tibetans
Here we have another of those rare cases of selective sweeps in humans: adaption to high altitude among Tibetans. Two genes have been found to produce this striking adaption to hypoxia and confirmed in this paper: EGLN1 and EPAS1. The authors suspect that other genes, notably ANGPT1, ECE1, and LEPR, are probably also involved in this adaption to such a demanding ecology.