 |
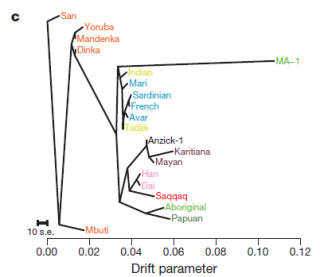
| Figure 4 | [c] (…) maximum likelihood tree. |
A recent study on the ancient DNA of human remains from Anzick (Montana, USA), dated to c. 12,500 calBP, confirms close ties to modern Native Americans, definitely discarding the far-fetched and outlandishly Eurocentric “Solutrean hypothesis” for the origins of Clovis culture (what pleases me greatly, I must admit).
While this fits well with the expectations (at least mine), there is some hidden data that has surprised me quite a bit: it sits at the bottom of a non-discussed formal test graph in which modern populations are compared with both Anzick and Tianyuan (c. 40,000 BP, North China). See below.
Morten Rasmussen et al., The genome of a Late Pleistocene human from a Clovis burial site in western Montana. Nature 2014. Pay per view → LINK [doi:10.1038/nature13025]
Abstract
Clovis, with its distinctive biface, blade and osseous technologies, is the oldest widespread archaeological complex defined in North America, dating from 11,100 to 10,700 14C years before present (bp) (13,000 to 12,600 calendar years bp)1, 2. Nearly 50 years of archaeological research point to the Clovis complex as having developed south of the North American ice sheets from an ancestral technology3. However, both the origins and the genetic legacy of the people who manufactured Clovis tools remain under debate. It is generally believed that these people ultimately derived from Asia and were directly related to contemporary Native Americans2. An alternative, Solutrean, hypothesis posits that the Clovis predecessors emigrated from southwestern Europe during the Last Glacial Maximum4. Here we report the genome sequence of a male infant (Anzick-1) recovered from the Anzick burial site in western Montana. The human bones date to 10,705 ± 35 14C years bp (approximately 12,707–12,556 calendar years bp) and were directly associated with Clovis tools. We sequenced the genome to an average depth of 14.4× and show that the gene flow from the Siberian Upper Palaeolithic Mal’ta population5 into Native American ancestors is also shared by the Anzick-1 individual and thus happened before 12,600 years bp. We also show that the Anzick-1 individual is more closely related to all indigenous American populations than to any other group. Our data are compatible with the hypothesis that Anzick-1 belonged to a population directly ancestral to many contemporary Native Americans. Finally, we find evidence of a deep divergence in Native American populations that predates the Anzick-1 individual.
Haploid DNA
The Y-DNA lineage of Anzick is Q1a2a1* (L54) to the exclusion of the common Native American subhaplogroup Q1a2a1a1 (M3). Among the modern compared sequences that of a Maya is the closest one.
The mtDNA belongs to the common Native American lineage D4h3a at its underived stage (root).
For starters I must explain that these underived haplotypes can only be found within mtDNA and never in modern Y-DNA (common misconception) because this one accumulates mutations every single generation, while the much shorter mtDNA does only occasionally. Hypothetically we could find the exact ancestor of some modern Y-DNA haplogroup in ancient remains but that would be like finding the proverbial needle in the haystack. On the other hand, finding the underived stage in mtDNA, be it ancient or modern, does not mean that we are before a direct ancestor but just a non-mutated relative of her, who can be very distant in fact.
Autosomal DNA
In this aspect, the Anzick man shows clearly strongest affinities to Native Americans, followed at some distance by Siberian peoples, particularly those near the Bering Strait.
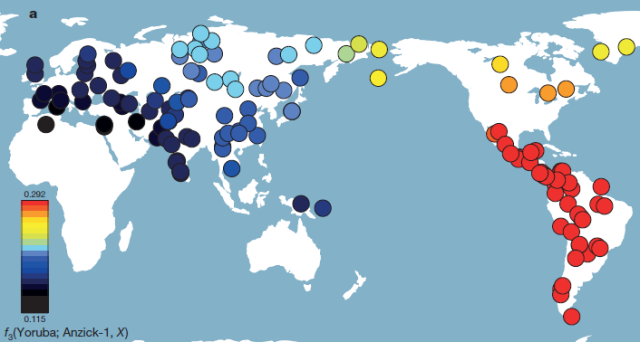 |
| Figure 2 | Genetic affinity of Anzick-1. a, Anzick-1 is most closely related to Native Americans. Heat map representing estimated outgroup f3-statistics for shared genetic history between the Anzick-1 individual and each of 143 contemporary human populations outside sub-Saharan Africa. (…) |
However Anzick-1 shows clearly closer affinity to the aboriginal peoples of Meso, Central and South America (collectively labeled as SA) and less so to those of Canada and the American Arctic (labeled as NA). No data was available from the USA.
This was pondered by the authors in several competing models of Native American ancestry:
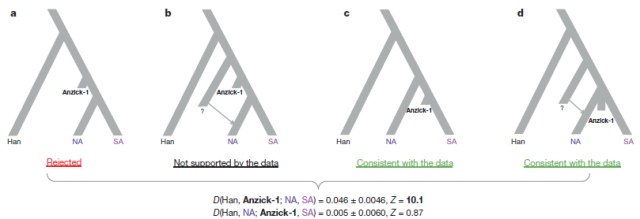 |
| Figure 3 | Simplified schematic of genetic models. Alternative models of the population history behind the closer shared ancestry of the Anzick-1 individual to Central and Southern American (SA) populations than Northern Native American (NA) populations; seemain text for further definition of populations. We find that the data are consistent with a simple tree-like model in which NA populations are historically basal to Anzick-1 and SA. We base this conclusion on two D-tests conducted on the Anzick-1 individual, NA and SA. We used Han Chinese as outgroup. a, We first tested the hypothesis that Anzick-1 is basal to both NA and SA populations using D(Han, Anzick-1; NA, SA). As in the results for each pairwise comparison between SA and NA populations (Extended Data Fig. 4), this hypothesis is rejected. b, Next, we tested D(Han, NA; Anzick-1, SA); if NA populations were a mixture of post-Anzick-1 and pre-Anzick-1 ancestry, we would expect to reject this topology. c, We found that a topology with NA populations basal to Anzick-1 and SA populations is consistent with the data. d, However, another alternative is that the Anzick-1 individual is from the time of the last common ancestral population of the Northern and Southern lineage, after which the Northern lineage received gene flow from a more basal lineage. |
The most plausible model they believe is “c”, in which Anzick-1 is close to the origin of the SA population, while NA diverged before him. However model “d” in which Anzick-1 is close to the overall Native American root but NA have received further inputs from a mystery population (presumably some Siberians, related to the Na-Dené and Inuit waves) is also consistent with the data. Choosing between both “consistent” models (or something in between) clearly requires further investigation.
Tianyuan and East Asian origins
All the above is very much within expectations, although refreshingly clarifying. But there is something in the formal tests (extended data fig. 5) that is most unexpected (but not discussed in the paper).
The formal f3 tests of ED-fig.5 a to e fall all within reasonable expectations. Maybe the most notable finding is that, after all, the pre-Inuit people of the Dorset culture (represented by the Saqqaq remains) left some legacy in Greenland, but they also show some extra affinity with several Siberian populations (notably the Naukan, Chukchi, Koryak and Yukaghir, in this order) before to any other Native Americans, including Aleuts).
But the really striking stuff is in figs.
f and
g, where it becomes obvious that the
Tianyuan remains of Northern China show not a tad of greater affinity to East Asians (nor to Native Americans) than to West Eurasians. Also two East Asian populations (Tujia and Oroqen) are considerably more distant than the bulk of East Asian peoples to Tianyuan but also to Aznick.
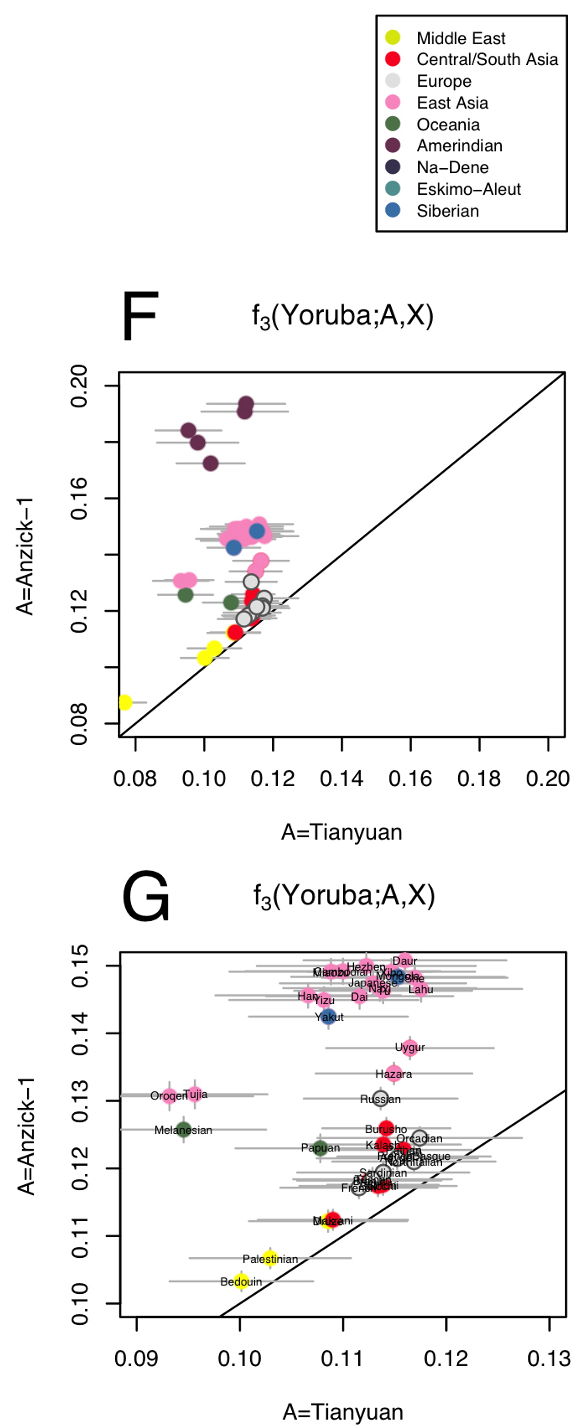 |
| Extended Data Figure 5 | Outgroup f3-statistics contrasted for different combinations of populations. (…) f, g, Shared genetic history with Anzick-1 compared to shared genetic history with the 40,000-year-old Tianyuan individual from China. |
This is very difficult to explain, more so as Tianyuan’s mtDNA haplogroup B4’5 is part of the East Asian and Native American genetic pool, and the authors make no attempt to do it.
The previous
study by Qiaomei Fu et al. (open access) placed Tianyuan’s autosomal DNA near the very root of Circum-Pacific populations (East Asians, Native Americans and Australasian Aborigines) but after divergence from West Eurasians:
 |
| From Qiaomei Fu 2013 |
They even had doubts about the position of Papuans (the only Australasian representation) in that tree, which they suspected an artifact of some sort.
Since I saw that graph (h/t to an anonymous commenter at Fennoscandian Ancestry) I am squeezing my brain trying to figure out a reasonable explanation, considering that the formal f3 test has almost certainly more weight than the ML tree made with an algorithm.
My first tentative explanation would be to imagine a shared triple-branch origin for Tianyuan, East Asians and West Eurasians, maybe c. 60 Ka ago (it must have been before the colonization of West Eurasia), to the exclusion of other, maybe isolated, ancient populations, whose admixture with the ancestors of the Tujia, Oroqen and Melanesians (maybe via Austronesians?) causes those striking low affinity values for these.
This would be a similar mechanism to the one explaining lower Tianyuan (and generally all ancient Eurasian) affinity for Palestinians (incl. Negev Bedouins) and also the Makrani, who have some African admixture and (in the Palestinian case) also, most likely, residual inputs from the remains of the first Out-of-Africa episode in Arabia.
However to this day we have no idea of which could be those hypothetical ancient isolated populations of East Asia. In normal comparisons such as ADMIXTURE analysis the Tujia and Oroqen appear totally normal within their geographic context, but this may be an artifact of not doing enough runs to reach higher K values, according to the cross-validation test, much more likely to discern the actual realistic components.
The matter certainly requires further research, which may well open new avenues for the understanding the genesis of Eurasian populations, particularly those from the East.











